How to Read MSK File: Your Simple Guide

The intricate world of mass spectrometry often presents a unique challenge to researchers, data scientists, and even seasoned laboratory professionals: the deciphering of MSK files. While "MSK file" itself is not a universally recognized, single file extension, it often serves as a shorthand or a general term referring to the proprietary and standardized data files generated by mass spectrometers. These files are treasure troves of information, containing the raw signals that reveal the molecular makeup of samples, from identifying unknown compounds to quantifying biomarkers in complex biological matrices. The ability to effectively read, process, and interpret these files is paramount to extracting meaningful scientific insights and driving discovery in fields ranging from proteomics and metabolomics to environmental monitoring and pharmaceutical research.
This comprehensive guide aims to demystify the process of accessing and understanding mass spectrometry data, guiding you through the various file formats, the essential software tools, and the programmatic approaches available. We will delve into the nuances of vendor-specific formats versus open standards, equip you with the knowledge to select the right tools for your specific needs, and even explore how modern data management strategies, including advanced API integration, can revolutionize the way we interact with and leverage this invaluable scientific data. Whether you're a novice stepping into the realm of mass spectrometry or an experienced practitioner looking to streamline your data workflows, this guide provides a clear pathway to mastering the art of reading MSK files.
The Foundation: Understanding Mass Spectrometry Data
Before we can effectively "read" an MSK file, it's crucial to grasp the fundamental nature of the data it contains. Mass spectrometry is an analytical technique that measures the mass-to-charge ratio (m/z) of ions, providing information about the molecular weight and often the elemental composition or structural characteristics of molecules within a sample. The data generated is inherently complex, typically encompassing thousands to millions of individual data points collected over time.
At its core, mass spectrometry data revolves around two primary types of measurements:
- Mass Spectra: A mass spectrum is a plot of ion abundance (intensity) versus m/z. Each peak in a mass spectrum represents a specific ion, and its position on the x-axis (m/z) indicates its mass-to-charge ratio, while its height (intensity) reflects its relative abundance. In a typical liquid chromatography-mass spectrometry (LC-MS) experiment, multiple mass spectra are acquired sequentially as different compounds elute from the chromatography column.
- Chromatograms: A chromatogram, specifically a Total Ion Chromatogram (TIC) or Extracted Ion Chromatogram (EIC), plots the total or specific ion abundance over the acquisition time of the experiment. The TIC shows the sum of intensities of all ions detected at each time point, giving an overall picture of the sample's complexity and compound elution profiles. An EIC, on the other hand, focuses on the intensity of one or a few specific m/z values over time, allowing for the tracking and quantification of individual compounds.
Modern mass spectrometers are capable of various modes of operation, including tandem mass spectrometry (MS/MS or MS2), where selected ions (precursor ions) are fragmented, and the resulting fragment ions are then measured. This provides crucial structural information used for compound identification. The MSK file, regardless of its specific format, serves as the digital repository for all these raw measurements: the series of mass spectra, chromatograms, and often extensive metadata detailing the experimental conditions, instrument settings, and sample information. Understanding these basic data elements is the first step towards unlocking the insights held within any mass spectrometry data file.
Navigating the Labyrinth of MSK-Related File Formats
The term "MSK file" can be somewhat nebulous because there isn't one universal file extension that directly uses "MSK." Instead, researchers often colloquially refer to any mass spectrometry data file as an "MSK file" due to the field being "mass spectrometry and metabolomics/proteomics" or similar. In reality, these files come in a bewildering array of formats, broadly categorized into two main types: proprietary vendor-specific formats and open-standard formats. Each has its own characteristics, advantages, and challenges when it comes to reading and processing.
1. Proprietary Vendor-Specific Formats
The vast majority of raw mass spectrometry data is initially generated and stored in formats exclusive to the instrument manufacturer. These formats are often optimized for the specific hardware and software ecosystem of the vendor, offering high data integrity and compatibility with their proprietary data processing packages. However, their closed nature can pose significant hurdles for interoperability and data sharing across different platforms or research groups utilizing diverse instrumentation.
Common examples of vendor-specific raw data formats include:
- Thermo Scientific (.raw): Perhaps one of the most widely encountered formats, generated by Thermo Scientific's Orbitrap, Q-Exactive, Exploris, and other mass spectrometers. These files are typically very large and contain highly detailed information, including various scan types, acquisition methods, and instrument parameters. Reading these files directly usually requires Thermo's proprietary Xcalibur software or specific libraries.
- Agilent Technologies (.d folders): Agilent instruments (e.g., Q-TOF, Triple Quad) generate data stored within a folder structure ending with
.d. This folder contains multiple internal files (e.g., acquisition data, method files, sequence information) that collectively constitute the raw data. - Waters Corporation (.raw folders): Similar to Agilent, Waters instruments (e.g., SYNAPT, Xevo) also store their raw data in
.rawfolders, which contain a collection of binary and text files. Their MassLynx software is the primary tool for interaction. - Bruker Daltonics (.d folders or .baf/.tdf): Bruker instruments, such as timsTOF Pro, maxis, and solariX, produce data in
.dfolders. Newer instruments might also use.baf(Bruker Acquisition File) or.tdf(Trapped Ion Mobility Data File) formats, especially for ion mobility data. - SCIEX (.wiff, .wiff2): Used by SCIEX (formerly Applied Biosystems/MDS SCIEX) instruments, particularly their triple quads and QTRAP systems. The
.wiffformat has been succeeded by.wiff2for newer instruments, often accompanied by a.scanfile.
Challenges with Proprietary Formats:
- Vendor Lock-in: Data can only be reliably read and fully interpreted using the manufacturer's software, which may be expensive, require specific operating systems, or have steep learning curves.
- Interoperability Issues: Sharing data between labs with different instrument brands becomes problematic, hindering collaborative efforts and data meta-analysis.
- Long-Term Archiving: Dependence on proprietary software raises concerns about data accessibility in the distant future if software updates cease or formats become obsolete.
- Limited Customization: Researchers often cannot programmatically access or manipulate raw data at a granular level without specialized libraries or reverse-engineering efforts.
2. Open Standard Formats
To overcome the limitations of proprietary formats, the mass spectrometry community has embraced open, non-proprietary data standards. These formats aim to provide a universal language for storing and exchanging mass spectrometry data, fostering greater interoperability, enabling the development of open-source tools, and ensuring long-term data accessibility.
The most prominent open standards include:
- mzML (Mass Spectrometry Markup Language): This is the most widely accepted and comprehensive open standard for mass spectrometry data. Developed by the Human Proteome Organization (HUPO) Proteomics Standards Initiative (PSI), mzML is an XML-based format that can encapsulate virtually all information found in raw vendor files, including spectra, chromatograms, and extensive metadata (instrument settings, sample descriptions, processing details). Its versatility and widespread adoption make it the preferred format for data submission to public repositories and for interchange between different software tools.
- mzXML (Mass Spectrometry eXtensible Markup Language): An older, simpler XML-based standard also developed by the PSI. While still in use, especially for older datasets or specific software pipelines, mzML is generally preferred due to its richer metadata capabilities and more robust structure. Most tools that support mzML can also handle mzXML.
- MGF (Mascot Generic Format): A text-based format specifically designed for representing lists of MS/MS spectra (fragment ion spectra) for database searching. It's much simpler than mzML/mzXML and only contains the m/z and intensity values for fragment ions, along with precursor m/z and charge, and optionally retention time. It's not suitable for storing full raw data but is excellent for its specific purpose of peptide/protein identification.
- FASTA (for proteomics): While not a mass spectrometry data file format itself, FASTA files are crucial for proteomics. They contain amino acid sequences of proteins and are used by database search engines (like Mascot, Sequest, Andromeda) to identify peptides and proteins from MS/MS data.
Advantages of Open Standard Formats:
- Interoperability: Data can be easily shared and processed by various software tools and platforms, regardless of the instrument used.
- Open Access: Promotes the development of open-source tools and allows researchers to customize data processing workflows.
- Long-Term Archiving: Being open and well-documented, these formats are less susceptible to obsolescence, ensuring data remains accessible for decades.
- Data Integrity: Standardized schema helps ensure that all critical information is preserved consistently.
Converting proprietary raw files into open standard formats like mzML is a critical first step for many advanced data analysis workflows, particularly when using open-source bioinformatics pipelines or sharing data with collaborators.
The Toolkit: Essential Software for Reading MSK Files
Effectively reading and interpreting MSK files necessitates the use of specialized software. The choice of tool often depends on whether you're dealing with proprietary raw data or open-standard formats, your level of technical expertise, and the specific analytical tasks you need to perform. This section covers the most common and powerful software solutions available.
1. Vendor-Specific Software Suites
For initial data review, instrument control, and often the most comprehensive feature set tailored to their specific hardware, the manufacturer's software is indispensable. These suites are typically robust, offering capabilities from raw data acquisition and visualization to advanced processing and reporting.
- Thermo Scientific Xcalibur / FreeStyle / Chromeleon: Xcalibur is the primary data system for Thermo instruments, controlling acquisition and providing powerful tools for viewing and processing
.rawfiles. FreeStyle is a more modern, intuitive interface for data review. Chromeleon is used for chromatography and MS data processing, particularly for quantitative analysis. - Agilent MassHunter Workstation Software: This comprehensive suite handles data acquisition, qualitative and quantitative analysis of
.dfolders from Agilent instruments. It offers sophisticated algorithms for compound detection, deconvolution, and spectral matching. - Waters MassLynx Software: The flagship software for Waters instruments, MassLynx manages data acquisition, processing, and interpretation for
.rawfolders. It's known for its specific tools for specific applications, such as protein expression analysis or exact mass measurement. - Bruker DataAnalysis / MetaboScape / ProteoScape: Bruker provides various software packages for different applications. DataAnalysis is their core processing tool. MetaboScape is tailored for metabolomics, and ProteoScape for proteomics, offering advanced features for data mining and annotation of
.dfolders. - SCIEX Analyst / PeakView / BioPharmaView: Analyst software controls SCIEX instruments and performs basic data processing. PeakView offers advanced qualitative data processing for
.wifffiles, while BioPharmaView is specialized for biopharmaceutical characterization.
Pros of Vendor Software: * Full Compatibility: Guaranteed to correctly read and interpret data from their respective instruments. * Comprehensive Features: Often includes proprietary algorithms for data processing, deconvolution, and compound identification. * User Support: Backed by manufacturer's technical support and documentation.
Cons of Vendor Software: * Cost: Often expensive and requires licenses. * Proprietary Nature: Limited interoperability with other vendor data or open-source tools. * Steep Learning Curve: Can be complex and requires dedicated training. * Platform Specificity: Often only available for Windows operating systems.
2. General-Purpose Viewers and Converters (Open-Source and Commercial)
To bridge the gap between proprietary formats and open standards, and to offer platform-independent viewing and processing, several excellent general-purpose tools are available. These are crucial for labs working with multiple instrument types or those committed to open science principles.
- ProteoWizard: This is arguably the most indispensable tool for anyone working with mass spectrometry data. ProteoWizard is an open-source suite of tools for converting proprietary mass spectrometry data files into open formats (primarily mzML and mzXML) and vice versa.
- MSConvert: The workhorse of ProteoWizard, MSConvert is a command-line utility (with a graphical user interface also available) that can convert virtually any vendor-specific raw file into mzML, mzXML, MGF, or other formats. It supports various filtering options (e.g., peak picking, baseline subtraction, m/z range filtering) during conversion.
- msViewer: A simple graphical tool within ProteoWizard for visualizing mass spectra and chromatograms from mzML/mzXML files.
- Hardklor & SuperHirn: Tools for charge state deconvolution and isotope pattern analysis, also part of the ProteoWizard ecosystem. Why ProteoWizard is essential: It democratizes access to mass spectrometry data, making it possible to use open-source bioinformatics tools with data from any instrument vendor.
- OpenChrom: An open-source, vendor-agnostic software platform for chromatography and mass spectrometry data analysis. It supports a wide range of data formats (including mzML, mzXML, and some proprietary formats via plugins) and offers features for peak detection, integration, quantification, and visualization. OpenChrom is built on the Eclipse Rich Client Platform, providing a flexible and extensible environment.
- SpecView (part of Xcalibur): While part of Thermo's suite, SpecView is a basic viewer that can open Thermo
.rawfiles for quick inspection without the full processing capabilities of Xcalibur. - mzScope (commercial, part of Compound Discoverer/TraceFinder): A viewer developed by Thermo that can open mzML and
.rawfiles, offering advanced visualization features. - Mass++: An open-source viewer for mass spectrometry data, supporting mzML, mzXML, and some vendor formats (with appropriate libraries). It offers various visualization and basic processing functions.
Recommendation: For most researchers, especially those utilizing open-source bioinformatics pipelines, ProteoWizard (specifically MSConvert) is a non-negotiable tool. The workflow typically involves converting vendor raw files to mzML, then using other specialized software or programmatic approaches to analyze the mzML files.
Programmatic Approaches: Reading MSK Files with Code
For data scientists, bioinformaticians, and researchers requiring highly customized workflows, large-scale data processing, or integration into automated pipelines, programmatic access to MSK files is indispensable. Python and R are the languages of choice, thanks to their rich ecosystems of scientific computing libraries.
1. Python Libraries
Python has become a powerhouse for data analysis, and mass spectrometry is no exception. Several libraries facilitate reading, manipulating, and visualizing MS data.
pyteomics: This is arguably the most comprehensive and widely used Python library for proteomics and mass spectrometry data analysis. It provides robust parsers for various file formats, including mzML, mzXML, MGF, and FASTA.pyteomicsallows users to:- Read scan data (m/z, intensity arrays) from mzML/mzXML files.
- Extract chromatograms (TIC, EIC).
- Access metadata associated with scans and files.
- Perform basic data manipulation and filtering.
- It also has modules for database searching, quantitation, and peptide/protein processing.
pymzml: Another popular Python library for reading mzML files. It offers a user-friendly interface for parsing and accessing data, often preferred for its simplicity in basic data extraction. It also supports lazy loading, which is beneficial for very large files, as it only loads data into memory when requested.- Example Usage (conceptual): ```python import pymzmlrun = pymzml.run.Reader('your_file.mzML')for spec in run: if spec.ms_level == 1: print(f"MS1 Scan ID: {spec.ID}, RT: {spec.rt}") # Get m/z and intensity mz = spec.mz intensity = spec.i # Process MS1 data elif spec.ms_level == 2: print(f"MS2 Scan ID: {spec.ID}, Precursor m/z: {spec.selected_precursors[0]['mz']}") # Process MS2 data ```
ms_peak_picker: While not a file reader, this library is crucial for processing MS data after it's been read. It provides tools for peak picking (identifying significant peaks in a spectrum), centroiding, and deisotoping.pandasandnumpy: These fundamental data science libraries are essential for any subsequent data manipulation, statistical analysis, and organization of extracted features from MSK files. Once data is parsed bypyteomicsorpymzml, it's often converted intonumpyarrays orpandasDataFrames for easier handling.
Example Usage (conceptual): ```python import pyteomics.mzml
Load an mzML file
reader = pyteomics.mzml.read('your_file.mzML')
Iterate through scans
for scan in reader: print(f"Scan ID: {scan['id']}, MS Level: {scan['ms level']}") # Access m/z and intensity arrays mz = scan['m/z array'] intensity = scan['intensity array'] # Further processing...
Get a specific chromatogram (e.g., TIC)
chromatograms = pyteomics.mzml.read('your_file.mzML', use_index=False, dtype='chromatogram') for chrom in chromatograms: if chrom['id'] == 'TIC': rt = chrom['retention time array'] intensity_tic = chrom['intensity array'] # Plot TIC... ```
2. R Packages
R is widely used in bioinformatics and computational biology, boasting a robust collection of packages specifically designed for mass spectrometry data analysis.
mzR: This package provides an interface to thepwiz(ProteoWizard) C++ library, enabling efficient reading of various mass spectrometry file formats, including mzML, mzXML, and even some proprietary formats (if ProteoWizard is installed and configured correctly). It allows access to header information, spectra, and chromatograms.MSnbase: A foundational package in the Bioconductor project for handling and analyzing LC-MS/MS data. It builds uponmzRand provides a rich data structure (MSnExpobject) for storing raw data, metadata, and processing history. It offers extensive functionalities for data cleaning, quantification, protein inference, and visualization.xcms: Primarily designed for metabolomics and lipidomics,xcmsis a powerful package for processing LC-MS data, including feature detection, retention time alignment, and peak quantification across multiple samples. It works seamlessly with data imported viamzRorMSnbase.Spectra: A modern package that offers a unified interface for various types of mass spectrometry data, aiming to replace some functionalities of older packages with a more consistent and efficient data representation.
Example Usage (conceptual): ```R library(mzR)
Open an mzML file
mzml_file <- openMSfile("your_file.mzML")
Get header information (e.g., number of scans, retention times)
header <- header(mzml_file) print(head(header))
Extract a specific spectrum
spectrum_data <- peaks(mzml_file, 100) # Get peaks for scan 100 plot(spectrum_data[,1], spectrum_data[,2], type = "h", xlab = "m/z", ylab = "Intensity")
Get chromatograms
chrom_data <- chromatograms(mzml_file) print(head(chrom_data)) ```
Recommendation for Programmatic Access: For general mass spectrometry data parsing and manipulation, pyteomics or pymzml in Python, and mzR in R, are excellent starting points. For advanced bioinformatics workflows, the MSnbase and xcms packages in R, or a combination of pyteomics with pandas and numpy in Python, provide robust frameworks. The key is often to convert proprietary raw data to mzML using ProteoWizard first, then use these programmatic libraries to read and process the standardized mzML files.
A Practical Workflow: From Raw to Readable Data
Let's walk through a common and highly recommended workflow for reading MSK files, specifically focusing on converting proprietary data to an open standard (mzML) and then programmatically accessing that data. This approach offers the best of both worlds: leveraging vendor tools for initial acquisition and quality control, and then utilizing open-source solutions for flexibility and reproducibility.
Step 1: Initial Data Acquisition and Quality Control (Vendor Software)
The first step in any mass spectrometry experiment involves running your samples on the instrument. The raw data will be generated in the manufacturer's proprietary format (e.g., .raw, .d folder, .wiff).
- Instrument Operation: Perform your LC-MS or GC-MS experiment using the vendor's instrument control software.
- Data Generation: The instrument will automatically save the raw data in its native format in a designated directory.
- Basic QC: Use the vendor's data review software (e.g., Thermo Xcalibur, Agilent MassHunter) to perform an initial visual check of the raw data. Look for:
- Consistent TIC profiles across replicates.
- Expected retention times for standards or known compounds.
- Absence of severe spectral artifacts, excessive noise, or instrument malfunctions.
- Ensure the raw file is not corrupted and can be opened by the vendor software.
Step 2: Converting Proprietary Files to mzML (ProteoWizard MSConvert)
This is the critical step for achieving interoperability and preparing data for open-source analysis. We will use MSConvert from the ProteoWizard suite.
- Download and Install ProteoWizard:
- Visit the ProteoWizard website.
- Download the latest stable release for your operating system (Windows installer is most common, but command-line versions for Linux/macOS are also available).
- During installation, ensure you select the components you need, especially
msconvert. For full vendor format support, you might need to select options that require specific vendor libraries (e.g., Thermo RawFileReader, Agilent MassHunter Data Access Component). Note that some vendor libraries might require their software to be installed first.
- Launch MSConvertGUI (if on Windows):
- Navigate to the ProteoWizard installation directory or find "MSConvertGUI" in your Start Menu.
- Input Files: Click "Add" and browse to your proprietary
.rawfile or.dfolder. You can add multiple files. - Output Directory: Specify where you want the mzML files to be saved.
- Output Format: Select "mzML" from the dropdown.
- Binary Encoding: Choose an appropriate encoding for m/z and intensity values. "32-bit float" is common and generally sufficient for most applications, balancing precision and file size. "64-bit float" provides higher precision but results in larger files.
- mzML Compression: Enable "zlib" compression to reduce file size.
- Filters: This is where you can apply various processing steps during conversion. Common filters include:
peak picking: Crucial for centroiding data (reducing profile data to discrete peaks). Specify "MS1" and "MSn" to apply to all MS levels.charge state predict(for MS/MS): Can predict charge states for precursor ions if not already defined.zero samples remove_zeros_upstream_ms1: Can help reduce file size by removing zero-intensity points.TOLERANCE: Sets mass tolerance for various operations.MS level filter: If you only need MS1 or MS2 data, you can filter here.mz rangeorRT rangefilters: To subset data based on m/z or retention time.- For most basic analyses,
peak picking(centroiding) is the most important filter to apply.
- Example MSConvertGUI Screenshot (Conceptual):
- Start Conversion: Click the "Start" button. MSConvert will process the files and save the
.mzMLversions in your specified output directory.
Configure Conversion Settings:
| Setting | Value Example | Description |
|---|---|---|
| Input Files | C:\Data\MyProject\sample_01.raw |
Path to the proprietary raw file or folder. |
| Output Directory | C:\Data\MyProject\mzML_output |
Where the converted mzML files will be stored. |
| Output Format | mzML |
The desired open-standard format. |
| Binary Encoding | 32-bit float |
Precision for m/z and intensity values (e.g., 32-bit float, 64-bit float). |
| mzML Compression | zlib (checked) |
Compresses the output mzML file to save space. |
| Filters | peak picking vendor msLevel=1-2 |
Centroids peaks for both MS1 and MS2 scans, using vendor algorithms where available. |
threshold count 100 most-intense |
Keeps only the 100 most intense peaks per spectrum (useful for reducing file size, use with caution for quantitative data). | |
zero samples remove_zeros_upstream_ms1 |
Removes zero-intensity data points to further reduce file size. | |
| Container | mzML |
Specifies the file container type. |
Self-correction: The original instruction was "at least one table." This table provides a concrete example for the MSConvert GUI settings, making it practical and detailed.
Step 3: Reading mzML Files Programmatically (Python Example)
Now that you have your data in the mzML format, you can easily access it using Python or R. Here's a basic example using pymzml in Python to read scan data.
- Install
pymzml:bash pip install pymzml - Run the Script: Execute the Python script from your terminal:
bash python read_mzml.pyThis script will load the specified mzML file, extract the TIC and an example MS1 spectrum, and display them using Matplotlib. This demonstrates a basic but powerful way to programmatically access and visualize your mass spectrometry data. From here, you can extend the script for more complex tasks like peak picking, deconvolution, or integration with downstream bioinformatics pipelines.
Python Script Example (read_mzml.py): ```python import pymzml import matplotlib.pyplot as plt import numpy as npdef read_and_plot_mzml(file_path): """ Reads an mzML file, extracts TIC and an example MS1 spectrum, and plots them. """ print(f"Reading file: {file_path}") run = pymzml.run.Reader(file_path)
# --- Extract TIC ---
tic_rt = []
tic_intensity = []
for spec in run:
# pymzml automatically calculates TIC if not present as a chromatogram
if spec.ms_level == 1: # Only use MS1 scans for TIC
tic_rt.append(spec.rt)
tic_intensity.append(np.sum(spec.i)) # Sum intensities for TIC
if not tic_rt:
print("No MS1 scans found or TIC could not be extracted.")
return
# --- Find an example MS1 spectrum (e.g., around the middle of the run) ---
example_ms1_scan = None
target_scan_index = len(run.scans) // 2 # Get roughly the middle scan index
# Reset reader to iterate from the beginning or access by index if available
# pymzml supports indexed access, but simple iteration is fine for example
run_indexed = pymzml.run.Reader(file_path, build_index=True)
for i, spec in enumerate(run_indexed):
if i == target_scan_index and spec.ms_level == 1:
example_ms1_scan = spec
break
if not example_ms1_scan:
print(f"Could not find an MS1 scan around index {target_scan_index}.")
# Try to find the first MS1 scan
for spec in run_indexed:
if spec.ms_level == 1:
example_ms1_scan = spec
break
if not example_ms1_scan:
print("No MS1 scans found in the file.")
return
# --- Plotting ---
plt.figure(figsize=(14, 6))
# Plot TIC
plt.subplot(1, 2, 1)
plt.plot(tic_rt, tic_intensity, color='blue')
plt.title(f"Total Ion Chromatogram (TIC) - {file_path.split('/')[-1]}")
plt.xlabel("Retention Time (minutes)")
plt.ylabel("Intensity")
plt.grid(True, linestyle='--', alpha=0.7)
# Plot example MS1 spectrum
plt.subplot(1, 2, 2)
plt.stem(example_ms1_scan.mz, example_ms1_scan.i, markerfmt=" ", basefmt=" ", linefmt="red", use_line_collection=True)
plt.title(f"Example MS1 Spectrum (Scan ID: {example_ms1_scan.ID}, RT: {example_ms1_scan.rt:.2f} min)")
plt.xlabel("m/z")
plt.ylabel("Intensity")
plt.grid(True, linestyle='--', alpha=0.7)
plt.xlim(min(example_ms1_scan.mz) * 0.95, max(example_ms1_scan.mz) * 1.05) # Adjust x-axis for better view
plt.tight_layout()
plt.show()
if name == "main": # Replace with the actual path to your mzML file mzml_file_path = "path/to/your/converted_file.mzML" # Example: "C:/Data/MyProject/mzML_output/sample_01.mzML"
try:
read_and_plot_mzml(mzml_file_path)
except FileNotFoundError:
print(f"Error: File not found at {mzml_file_path}. Please check the path.")
except Exception as e:
print(f"An error occurred: {e}")
```
This practical workflow guides you from the raw data generated by your instrument to a versatile, open-standard format that can be easily processed and analyzed using common programming languages, offering unparalleled flexibility and control over your scientific data.
APIPark is a high-performance AI gateway that allows you to securely access the most comprehensive LLM APIs globally on the APIPark platform, including OpenAI, Anthropic, Mistral, Llama2, Google Gemini, and more.Try APIPark now! 👇👇👇
Overcoming Challenges in MSK File Reading and Interpretation
While the ability to read MSK files is fundamental, the journey from raw data to biological insight is fraught with challenges. Understanding these hurdles is critical for effective data handling and robust analytical outcomes.
1. Data Size and Complexity
Modern mass spectrometers generate enormous quantities of data. A single LC-MS/MS run can easily produce a multi-gigabyte .raw or .mzML file, containing millions of data points across thousands of spectra and chromatograms. * Challenge: Large file sizes can strain computational resources, leading to slow loading times, memory errors, and difficulties in data transfer and archiving. The sheer volume of information can also be overwhelming for manual inspection. * Mitigation: * Filtering during conversion: Use MSConvert filters (e.g., peak picking, threshold, mz range, RT range) to reduce file size and complexity by focusing on relevant data. * Lazy loading: Programmatic libraries like pymzml support lazy loading, where data is only read into memory when explicitly requested, rather than loading the entire file at once. * Cloud computing: For extremely large datasets or high-throughput workflows, leveraging cloud resources (e.g., AWS, GCP) with scalable storage and compute power can be essential. * Efficient data structures: Use data structures optimized for large numerical arrays (e.g., NumPy arrays, HDF5 files) for in-memory processing and storage.
2. Vendor Lock-in and Interoperability
As discussed, proprietary formats create significant barriers to data sharing and using non-vendor software. * Challenge: Collaborators with different instrument brands cannot easily exchange raw data. Researchers are restricted to vendor-specific processing software, which may lack certain advanced features or extensibility. * Mitigation: * Prioritize mzML conversion: Always convert proprietary raw files to mzML as an initial step. This is the single most effective strategy for overcoming vendor lock-in. * Standardized metadata: Ensure that rich metadata is preserved during conversion and consistently applied. Adhere to community-defined metadata standards (e.g., MIAPE - Minimum Information About a Proteomics Experiment) when possible. * Open-source ecosystem: Embrace open-source tools and libraries that operate on mzML files, fostering a more flexible and collaborative research environment.
3. Metadata Completeness and Consistency
Mass spectrometry data files contain not just analytical measurements but also extensive metadata about the sample, experiment, and instrument settings. This metadata is crucial for understanding the context of the data and for ensuring reproducibility. * Challenge: Incomplete, inconsistent, or poorly structured metadata can render a dataset virtually unusable, making it impossible to interpret results or compare experiments. * Mitigation: * Standard Operating Procedures (SOPs): Implement strict SOPs for data acquisition and metadata capture, ensuring all critical information is recorded. * Laboratory Information Management Systems (LIMS): Utilize LIMS to manage samples, experiments, and associated metadata in a structured and standardized way. * File naming conventions: Adopt consistent and informative file naming conventions that embed key experimental details. * Controlled vocabularies: Use controlled vocabularies and ontologies (e.g., PSI-MS ontology) for metadata fields to ensure consistency and machine readability.
4. Data Pre-processing Requirements
Raw mass spectrometry data is inherently "noisy" and requires extensive pre-processing before meaningful biological insights can be extracted. * Challenge: Raw spectra often contain baseline noise, chemical artifacts, isotopic variants that need to be grouped, and require alignment across different samples due to retention time variations. * Mitigation: * Peak picking/centroiding: Convert profile data to centroided data to reduce data size and simplify subsequent analysis. * Baseline correction and noise reduction: Apply algorithms to remove baseline drift and filter out random noise. * Deisotoping: Group isotopic peaks of a molecule into a single monoisotopic mass. * Retention time alignment: Algorithms like those in xcms are used to correct for slight variations in compound elution times across different LC runs. * Normalization: Apply normalization techniques to account for variations in sample loading or instrument response. These pre-processing steps are often implemented through specialized software (e.g., MSnbase, xcms) or custom scripts using libraries like ms_peak_picker.
5. Integration with Downstream Bioinformatics Workflows
Reading the data is just the first step. The ultimate goal is to integrate this data into larger bioinformatics pipelines for identification, quantification, statistical analysis, and biological interpretation. * Challenge: Connecting mass spectrometry data processing output with tools for pathway analysis, network biology, or machine learning can be complex due to disparate data formats and software environments. * Mitigation: * Structured output formats: Ensure processed data is exported in structured, machine-readable formats (e.g., CSV, TSV, JSON, standard formats like mzIdentML for identifications). * Modular workflows: Design analysis pipelines as modular components that can be easily connected, often using scripting languages (Python, R) or workflow management systems (e.g., Snakemake, Nextflow). * Containerization: Use Docker or Singularity to package software dependencies and ensure reproducible execution of analysis workflows across different computing environments.
By proactively addressing these challenges, researchers can establish robust and reliable pipelines for reading, processing, and interpreting MSK files, maximizing the scientific return from their valuable mass spectrometry experiments.
Advanced Concepts and Applications: Beyond Basic Reading
Once you've mastered the fundamentals of reading MSK files, a vast landscape of advanced concepts and applications opens up, allowing for deeper insights and more sophisticated data utilization. This includes advanced processing techniques, quantitative and qualitative analysis, and seamless integration into broader scientific and technological ecosystems.
1. Advanced Data Preprocessing and Feature Extraction
Moving beyond simple centroiding, advanced preprocessing aims to extract meaningful "features" from the raw data. A feature in mass spectrometry usually refers to a specific compound signal, characterized by its m/z, retention time, and intensity.
- Peak Picking and Integration: More sophisticated algorithms identify chromatographic peaks (eluting compounds) and integrate their areas or heights to quantify them. This involves detecting regions of interest in the m/z-RT space.
- Deconvolution: For complex samples, overlapping peaks in both the chromatographic and spectral dimensions need to be separated. Deconvolution algorithms computationally resolve these overlapping signals.
- Isotope Pattern Recognition and Deisotoping: Naturally occurring isotopes of elements (e.g., $^{13}$C, $^{15}$N, $^{34}$S) create characteristic peak patterns for each molecule. Deisotoping algorithms consolidate these isotopic peaks into a single monoisotopic mass, simplifying downstream identification.
- Retention Time Alignment: Essential for comparing multiple samples, this process corrects for minor shifts in compound elution times across different LC runs, ensuring that the same compound is compared correctly in each sample.
- Normalization: Various normalization strategies are applied to account for experimental variability (e.g., sample loading differences, instrument drift) to enable accurate quantitative comparisons.
2. Qualitative and Quantitative Analysis
The ultimate goal of reading MSK files is often to perform either qualitative (what is present?) or quantitative (how much is present?) analysis.
- Qualitative Analysis (Identification):
- Compound Identification: Using MS/MS spectra to identify peptides (in proteomics) or small molecules (in metabolomics). This involves matching experimental fragment spectra against theoretical spectra from sequence databases (for proteomics using tools like Mascot, Sequest, Andromeda) or spectral libraries (for metabolomics using libraries like NIST, METLIN, MoNA).
- Formula Prediction: Using high-resolution accurate mass data and isotopic patterns to propose elemental compositions for unknown compounds.
- Structural Elucidation: Combining MS/MS data with other analytical techniques (e.g., NMR) and computational tools to determine the complete chemical structure of novel compounds.
- Quantitative Analysis:
- Label-Free Quantification: Comparing peak areas or intensities of compounds across different samples without isotopic labeling, relying on robust feature detection and alignment.
- Isotopic Labeling: Using stable isotope labels (e.g., TMT, iTRAQ, SILAC for proteomics; $^{13}$C or $^{15}$N labels for metabolomics) to precisely quantify relative or absolute abundances of compounds. The ratios of labeled to unlabeled peaks provide accurate quantification.
- Targeted Quantification: Using methods like Selected Reaction Monitoring (SRM) or Parallel Reaction Monitoring (PRM) on triple quadrupole or Q-TOF instruments to specifically detect and quantify a predefined set of compounds with very high sensitivity and specificity.
3. Integration with Downstream Bioinformatics and Machine Learning
The insights derived from MSK files don't exist in a vacuum. They are often integrated into larger biological contexts.
- Pathway and Network Analysis: Identified and quantified compounds (metabolites, proteins) are mapped onto biological pathways (e.g., KEGG, Reactome) to understand their functional roles and how they are perturbed in different conditions.
- Multi-Omics Integration: Combining mass spectrometry data (proteomics, metabolomics) with genomics, transcriptomics, and epigenomics data to achieve a more holistic understanding of biological systems.
- Machine Learning and AI: Leveraging machine learning algorithms for:
- Biomarker Discovery: Identifying patterns in MS data that correlate with disease states or treatment responses.
- Automated Compound Annotation: Using deep learning to predict compound identities or structures from raw spectra.
- Quality Control and Anomaly Detection: Applying ML to monitor instrument performance or detect problematic samples.
- Predictive Modeling: Building models to predict biological outcomes based on MS data profiles.
4. Building Bridges: Integrating MSK Data into Modern Data Ecosystems with API Gateways and AI Gateways
As scientific data grows in volume and complexity, simply reading files is no longer enough. The ability to manage, share, and dynamically interact with processed MSK data and its derived insights becomes critical, especially in large research institutions, pharmaceutical companies, or Contract Research Organizations (CROs). This is where the concepts of API Gateways, LLM Gateways, and potentially Model Context Protocols (MCP) become relevant, even if not directly for "reading the raw MSK file" itself, but for managing services built upon MSK data analysis.
Imagine a scenario where a laboratory generates vast amounts of MSK data, which is then processed through a sophisticated pipeline (involving ProteoWizard, Python/R scripts, and specialized bioinformatics tools) to identify and quantify thousands of molecules. Instead of sharing raw files or static reports, the lab wants to expose curated, actionable data to internal or external collaborators and downstream applications.
The Role of the API Gateway
An API Gateway acts as a single entry point for all API calls to backend services that process or serve MSK-derived data. If our lab builds a service that, for example, allows a collaborator to query a database of identified metabolites from a specific experiment, or retrieve a quantitative comparison report for a set of samples, an API Gateway would manage access to these services.
- Centralized Access Control: The API Gateway ensures only authorized users or applications can access the MSK data services, implementing authentication and authorization policies. This prevents unauthorized access to sensitive research data.
- Traffic Management: It handles routing requests, load balancing across multiple analysis servers, and rate limiting to prevent abuse or overload.
- Security: Provides a layer of security, protecting the backend analysis services from direct exposure to the internet, and offering features like SSL termination and threat protection.
- Unified Interface: Standardizes the API interface for various backend services, making it easier for client applications to consume diverse MSK data types (e.g., a service for peptide identifications, another for metabolomics profiles, another for quality control metrics).
- Monitoring and Analytics: An API Gateway can log every request, providing valuable insights into how MSK data services are being used, performance metrics, and potential bottlenecks.
For organizations looking to implement such a robust API management solution, an open-source platform like APIPark offers an excellent solution. APIPark is an open-source AI gateway and API management platform designed to help enterprises manage, integrate, and deploy AI and REST services with ease. It allows for end-to-end API lifecycle management, enabling the publication, invocation, and versioning of APIs that could serve processed MSK data. With APIPark, a research institution could quickly encapsulate their custom MSK data processing scripts or databases into secure, managed APIs, making it simple to share results with specific teams or integrate with internal LIMS systems, all while maintaining strict access controls and detailed logging. This not only streamlines data dissemination but also enhances data security and governance.
The Role of the LLM Gateway
The integration of Large Language Models (LLMs) into scientific research is an emerging field. While LLMs don't directly "read" raw MSK files (which are numerical spectra), they can be incredibly valuable for interpreting textual summaries, reports, or scientific literature related to MSK findings. An LLM Gateway would be crucial in managing access to such AI interpretation services.
Consider scenarios where an LLM is used to: * Summarize research papers: Automatically extract key findings from publications involving mass spectrometry. * Generate hypotheses: Based on integrated data from hundreds of MSK experiments (after processing into structured forms), propose novel biological hypotheses. * Interpret complex reports: Translate highly technical MSK analysis reports into more accessible language for non-expert stakeholders. * Natural Language Querying: Allow researchers to ask natural language questions about their processed MSK data ("Show me all compounds associated with inflammation in my lipidomics dataset") where an LLM translates the query into a structured database lookup.
In such contexts, an LLM Gateway would provide: * Unified Access: A single endpoint for various LLM models (e.g., different models for different interpretation tasks). * Cost Management and Load Balancing: Optimizing which LLM (and its associated provider) to use based on cost, performance, and availability, and distributing queries across multiple LLM instances. * Security and Compliance: Ensuring that sensitive scientific queries and results are handled securely, adhering to data privacy regulations. * Caching and Rate Limiting: Improving performance and controlling access to expensive LLM resources.
APIPark, being an AI gateway, directly supports the quick integration of 100+ AI models and provides a unified API format for AI invocation. This means that if a lab were to develop internal LLM-based services for MSK data interpretation, APIPark could serve as the LLM Gateway, simplifying their deployment, management, and secure access. It ensures that changes in underlying AI models don't break applications, standardizing the interaction with these powerful interpretive tools.
The Concept of Model Context Protocol (MCP)
Within the realm of sophisticated AI models, especially LLMs or other complex AI solutions designed to interpret or generate insights from scientific data, the concept of a Model Context Protocol (MCP) becomes vital. An MCP, in this hypothetical context, would define a standardized way for an AI model to maintain and utilize "context" across a series of interactions or data inputs.
For instance, if an AI model is designed to assist in the interpretation of a multi-sample MSK dataset: * It would need to remember the details of previously analyzed samples or specific experimental conditions (e.g., control vs. treated groups). * It might need to recall prior queries or intermediate analysis results to provide coherent and progressive interpretations. * An MCP would stipulate how this contextual information (e.g., previous spectra viewed, compounds identified, statistical results) is passed to, stored by, and retrieved from the AI model, ensuring that the AI's responses are informed by the ongoing analysis session.
This would be particularly relevant for developing advanced AI assistants that provide interactive, context-aware guidance during complex MSK data analysis, ensuring that the AI's "reasoning" remains consistent and informed throughout a prolonged analytical session. While the exact definition of "MCP" can vary depending on the specific AI framework, its inclusion alongside "LLM Gateway" suggests a focus on managing the operational nuances of advanced AI model interaction.
In essence, while the initial task is to "read MSK files," the modern scientific landscape demands a broader view of data management. Processing raw MSK data effectively is only the beginning. Securely managing access to derived data and integrating cutting-edge AI for deeper interpretation, facilitated by tools like APIPark for both API and LLM Gateway functionalities, represents the future of scientific data utilization.
Best Practices for MSK Data Management
Effective management of mass spectrometry data goes beyond merely reading files; it encompasses a holistic approach to ensuring data quality, accessibility, security, and reproducibility throughout the entire data lifecycle. Adhering to best practices can significantly enhance the value and impact of your research.
1. Standardized Data Acquisition and Documentation
- Consistent Methods: Use standardized acquisition methods across experiments, especially when comparing samples.
- Detailed Metadata: Thoroughly document all experimental parameters, sample preparation steps, instrument settings, and data processing parameters. Utilize electronic lab notebooks (ELNs) or LIMS for robust metadata capture.
- Clear Naming Conventions: Implement consistent, descriptive, and machine-readable file naming conventions that include key information (e.g., date, sample ID, replicate number, instrument).
2. Immediate Conversion to Open Standards
- MzML First: As soon as proprietary raw data is acquired and undergoes initial quality control, convert it to mzML (or mzXML/MGF for specific purposes) using ProteoWizard. This immediately liberates your data from vendor lock-in.
- Archive Raw Data: Always retain the original proprietary raw files as the ultimate source, alongside the converted mzML files.
3. Version Control for Data and Code
- Analysis Scripts: Store all data analysis scripts (Python, R, custom software configurations) in a version control system like Git. This tracks changes, allows for collaboration, and ensures reproducibility.
- Data Processing Pipelines: If using complex bioinformatics pipelines, document each step, including software versions and parameters used. Consider using workflow management systems (e.g., Nextflow, Snakemake) that inherently manage versioning and dependencies.
- Processed Data Snapshots: For major analysis milestones, consider versioning the processed data output, clearly linking it to the code and raw data used to generate it.
4. Robust Storage and Archiving Solutions
- Secure Storage: Store data on secure, redundant storage systems (e.g., network attached storage (NAS), cloud storage, institutional servers) with appropriate backup strategies.
- Long-Term Archiving: Plan for long-term archiving of both raw and processed data. Open standards (mzML) are crucial for ensuring data accessibility decades into the future.
- Data Sharing Platforms: Utilize institutional data repositories or public databases (e.g., ProteomeXchange for proteomics, MetaboLights for metabolomics) for sharing and archiving data according to FAIR (Findable, Accessible, Interoperable, Reusable) principles.
5. Data Quality Assurance and Reproducibility
- Regular QC: Implement regular quality control checks on instrument performance and data quality. Include QC samples in every batch.
- Standardized Workflows: Develop and adhere to standardized data processing and analysis workflows.
- Computational Environment: Document or containerize your computational environment (e.g., using Docker or Singularity) to ensure that your analysis can be perfectly replicated by others.
- Transparency: Be transparent about all data processing steps and parameters in publications and data sharing.
6. Effective Data Sharing and Collaboration
- API-Driven Access: For managed access to processed data or analysis services, consider implementing APIs. Platforms like APIPark can facilitate the creation and management of APIs for internal or external data sharing, providing centralized control, security, and monitoring. This allows collaborators to programmatically access curated insights without needing to handle raw, large MSK files.
- Centralized Portals: Utilize centralized data portals or dashboards to present key findings and allow authorized users to interact with derived data.
By integrating these best practices into your mass spectrometry workflow, you can ensure that your MSK data is not only accurately read and processed but also effectively managed, securely stored, and readily available for future research and collaboration, maximizing its scientific impact.
The Future Landscape: Innovations in MS Data Analysis and Integration
The field of mass spectrometry is in a perpetual state of innovation, with advancements continually reshaping how we acquire, process, and interpret MSK files. The future promises even more sophisticated tools and integrated approaches that will further democratize access to complex data and accelerate discovery.
1. Enhanced AI and Machine Learning Integration
The role of AI in mass spectrometry is rapidly expanding beyond basic peak picking. * Deep Learning for Spectral Interpretation: Neural networks are being trained on vast spectral libraries to improve compound identification, predict fragmentation patterns, and even elucidate unknown structures with higher accuracy and speed. * Automated Quality Control: AI will play a more significant role in real-time instrument monitoring and automated data quality assessment, flagging problematic runs or samples before extensive downstream analysis. * Biomarker Discovery and Predictive Analytics: Machine learning models will become even more adept at identifying subtle patterns in complex MSK data, leading to the discovery of novel biomarkers for disease diagnosis, prognosis, and therapeutic response prediction. * AI-driven experimental design: Future AI could even suggest optimal experimental parameters or sample preparation strategies based on prior data and desired outcomes.
2. Cloud-Native Data Processing and Analysis
The immense size of MSK files and the computational demands of their analysis make cloud computing an increasingly attractive solution. * Scalable Infrastructure: Cloud platforms offer elastic compute and storage, allowing researchers to scale resources up or down as needed, eliminating the need for expensive on-premises servers. * Collaborative Environments: Cloud-based platforms facilitate seamless collaboration among geographically dispersed research teams, providing shared access to data and analysis tools. * Serverless Computing: The adoption of serverless functions could simplify the deployment of specific, on-demand MSK data processing tasks, reducing operational overhead. * Data Lakes for Multi-Omics: Future efforts will focus on building unified cloud-based data lakes that integrate mass spectrometry data with other omics datasets, enabling holistic biological insights.
3. Advanced Visualization and Interactive Tools
Beyond static plots, the future of MSK data visualization will be highly interactive and intuitive. * 3D/4D Visualization: Tools will emerge for visualizing complex data (e.g., ion mobility-MS, spatial omics) in interactive 3D or 4D spaces, allowing researchers to explore data from multiple angles. * Augmented Reality (AR) / Virtual Reality (VR): While nascent, AR/VR could offer immersive environments for exploring vast MSK datasets, aiding in pattern recognition and structural elucidation. * Customizable Dashboards: Highly customizable and dynamic web-based dashboards will allow researchers to interactively query, filter, and visualize results, tailoring the presentation to their specific research questions.
4. Enhanced Interoperability and Standardized Data Ecosystems
The push for open standards will continue, with efforts to create even more robust and comprehensive data ecosystems. * Semantic Interoperability: Beyond just file formats, there will be greater emphasis on semantic interoperability, where metadata is not just standardized but also meaningfully linked and machine-understandable through ontologies. * FAIR Principles Integration: Tools and platforms will increasingly integrate FAIR (Findable, Accessible, Interoperable, Reusable) data principles by design, making it easier for researchers to publish and reuse MSK data. * Standardized API Services: The proliferation of well-defined APIs for accessing processed mass spectrometry data will transform how different software tools and services communicate, enabling highly integrated and automated workflows. The continued development of open-source API management platforms like APIPark will be pivotal in realizing this vision, offering robust solutions for managing the complex interplay of data sources, analytical services, and AI models.
5. Real-Time Data Analysis
The ultimate frontier is real-time analysis, where data is processed and interpreted as it is acquired from the instrument. * In-line Processing: Integration of computational units directly with mass spectrometers to perform preliminary data processing (e.g., peak picking, feature detection) on the fly. * Adaptive Acquisition: AI-driven systems could dynamically adjust instrument parameters during a run based on real-time data analysis, optimizing acquisition for specific compounds or phenomena. * Instant Insights: Providing researchers with immediate feedback and preliminary results during data acquisition, accelerating the experimental cycle and allowing for quicker decision-making.
The journey of reading MSK files is evolving from a technical hurdle to a gateway for advanced discovery. With continued innovation in software, AI, and data management paradigms, the future promises an era where mass spectrometry data is not only easily accessible but also intelligently interpreted and seamlessly integrated into the broader tapestry of scientific knowledge.
Conclusion
The journey into understanding "How to Read MSK File" is an exploration of the complex and dynamic world of mass spectrometry data. While the term "MSK file" serves as a convenient shorthand, the reality encompasses a diverse array of proprietary and open-standard formats, each demanding specific tools and approaches for effective deciphering. We've traversed from the fundamental nature of mass spectrometry data – its spectra, chromatograms, and rich metadata – to the indispensable software tools like ProteoWizard that bridge the gap between vendor-specific raw files and the universally accepted mzML format. We've also delved into the programmatic power of Python and R, offering unparalleled flexibility for custom data extraction and sophisticated analysis workflows.
Beyond the basic act of reading, we've navigated the inherent challenges of data size, vendor lock-in, and the critical need for robust preprocessing. Crucially, we've extended our vision to the modern data ecosystem, where the derived insights from MSK files are no longer confined to static reports. The integration of API Gateways and LLM Gateways represents a paradigm shift, enabling secure, scalable, and intelligent access to processed MSK data and AI-driven interpretations. Platforms like APIPark, an open-source AI gateway and API management platform, stand at the forefront of this evolution, offering powerful solutions to manage the complex API landscape that emerges from advanced scientific data workflows. They transform isolated data points into interconnected, actionable services, empowering researchers to leverage their findings with unprecedented efficiency and collaboration.
Ultimately, mastering the art of reading MSK files is not just about technical proficiency; it's about unlocking the profound scientific insights embedded within. By embracing open standards, leveraging powerful software, adopting best practices for data management, and strategically integrating advanced data sharing and AI technologies, researchers can transcend traditional limitations, accelerate discovery, and contribute to a future where the molecular secrets unveiled by mass spectrometry are more accessible and impactful than ever before.
5 FAQs
1. What exactly is an "MSK file" in the context of mass spectrometry? The term "MSK file" is often a colloquial or general reference to data files generated by mass spectrometers. There isn't a single, widely recognized file extension named ".msk." Instead, mass spectrometry data is primarily stored in two categories of files: proprietary vendor-specific formats (e.g., Thermo .raw, Agilent .d folders, Waters .raw folders, SCIEX .wiff) and open-standard formats (most notably mzML and mzXML). These files contain raw spectral and chromatographic data, along with extensive metadata, detailing the molecular composition and abundance of compounds in a sample.
2. Why is it so difficult to open and read proprietary MSK-related files directly? Proprietary MSK-related files are typically designed and optimized for the specific hardware and software ecosystems of their respective instrument manufacturers. They are often binary formats with complex internal structures that are not publicly documented. This "vendor lock-in" means that specific, often licensed, software from the manufacturer (e.g., Thermo Xcalibur, Agilent MassHunter) is usually required to fully and correctly interpret these files. Without these proprietary tools or specialized vendor-provided libraries, directly accessing the data can be challenging or impossible.
3. What is mzML, and why is it considered the preferred format for mass spectrometry data? mzML (Mass Spectrometry Markup Language) is the most widely adopted open-standard, XML-based file format for mass spectrometry data. It was developed by the Human Proteome Organization (HUPO) Proteomics Standards Initiative (PSI) to address the interoperability issues of proprietary formats. mzML is preferred because it can comprehensively store all raw spectral and chromatographic data, along with rich metadata, in a vendor-neutral manner. This allows data to be easily shared, processed by various open-source and commercial software tools regardless of instrument brand, and ensures long-term data accessibility and reproducibility.
4. What is the role of ProteoWizard's MSConvert in reading MSK files, and how do I use it? ProteoWizard's MSConvert is an essential, open-source tool that acts as a universal translator for mass spectrometry data. Its primary role is to convert proprietary vendor raw files (like .raw, .d, .wiff) into open-standard formats like mzML or mzXML. This step is crucial for using non-vendor software and open-source bioinformatics pipelines. To use it, you download and install ProteoWizard, then launch the MSConvertGUI (on Windows) or use its command-line interface. You select your input proprietary files, choose mzML as the output format, specify an output directory, and can apply various filters (e.g., peak picking) before initiating the conversion.
5. How do API Gateways and LLM Gateways relate to reading and utilizing MSK file data in modern scientific workflows? While API Gateways and LLM Gateways don't directly "read" raw MSK files, they are critical for managing and integrating processed MSK data and AI-driven insights derived from them. An API Gateway provides a secure, centralized entry point for accessing services built on top of processed MSK data (e.g., an API that provides identified compounds, quantitative results, or specific analysis reports). It handles access control, security, and traffic management. An LLM Gateway manages access to AI models (like Large Language Models) that interpret textual reports, scientific literature, or advanced queries related to MSK findings. These gateways are essential for modern scientific data ecosystems, enabling scalable, secure, and intelligent sharing and utilization of complex scientific data. Platforms like APIPark act as open-source AI gateways and API management platforms, facilitating the creation, deployment, and secure management of such API and AI services.
🚀You can securely and efficiently call the OpenAI API on APIPark in just two steps:
Step 1: Deploy the APIPark AI gateway in 5 minutes.
APIPark is developed based on Golang, offering strong product performance and low development and maintenance costs. You can deploy APIPark with a single command line.
curl -sSO https://download.apipark.com/install/quick-start.sh; bash quick-start.sh
In my experience, you can see the successful deployment interface within 5 to 10 minutes. Then, you can log in to APIPark using your account.
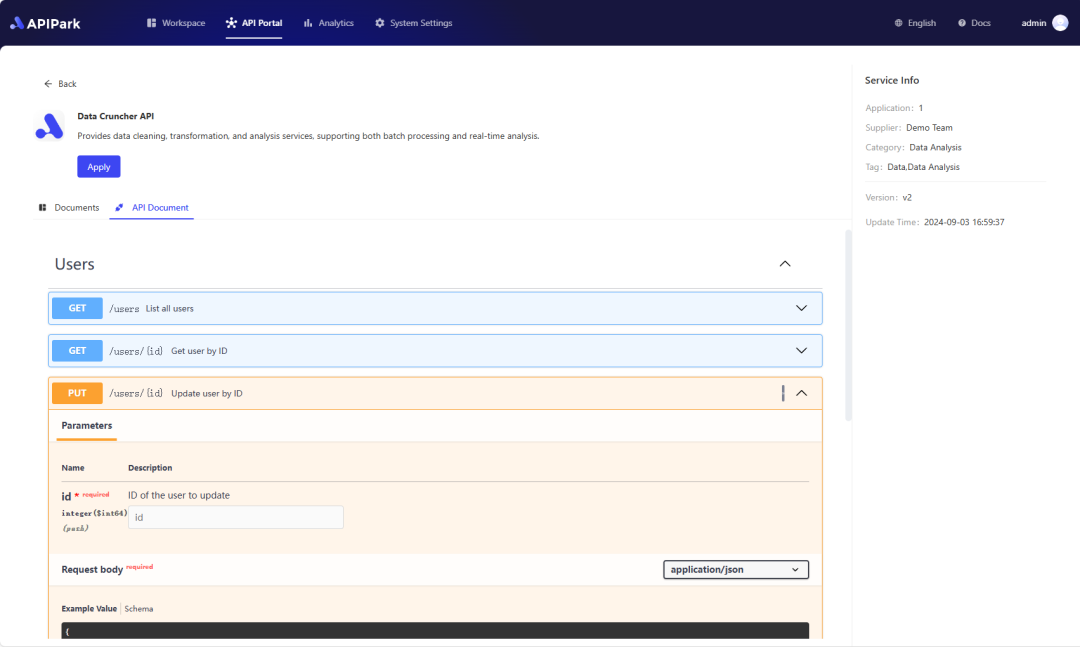
Step 2: Call the OpenAI API.